Cycle Sequencing
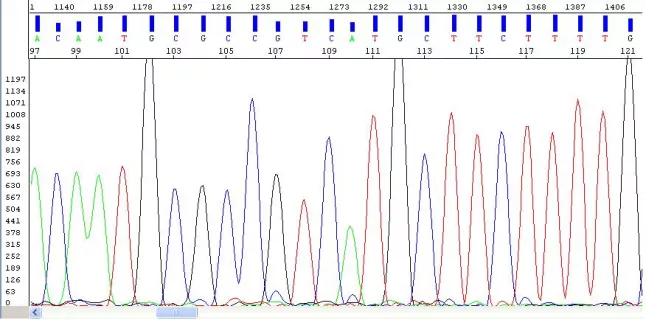
We use Applied Biosystem's Big Dye Terminator v3.1 Cycle Sequencing Kit for our sequencing reactions. PCR is employed to generate DNA fragments of varying lengths. Dye terminators stop the extension process, resulting in products with labelled ends. These are sorted by size and a laser is used to detect the labelled base at the terminated end. The resulting electropherogram is used to determine the sequence.
This video does a great job of explaining Cycle Sequencing.
Sequencing Primers
The manual for the Big Dye v3.1 kit provides several tips for creating good sequencing primers. These include:
- a length of at least 18 bases
- avoid repeats
- G/C content at 30-80%
- Tm above 45oC
- avoid secondary structures
Analyzing Sequencing Results
Results will be emailed to you. There are several freeware programs available. You can download Sequence Scanner from Applied Biosystems.
Alternately, you can email or make an appointment with the Genetics Lab and we can go over any questions you may have.
Fragment Analysis
The ABI 3130xl is capable of detecting five dyes per well, one of which being the LIZ size standard. The matrix standards that are needed to detect particular dye combinations are illustrated on this Dye Set Card. The DS-33 set is the one used by the Genetics Lab.
Applied Biosytems has a comprehensive guide about DNA Fragment Analysis using their system.
Our facility uses GeneScanner. Your data can also be analyzed with PeakScanner, which can be downloaded here.
If you would like to learn more about the different types of fragment analysis read this document about Fragment Sizing.